
Out of all the interesting methodologies I have encountered in the first 18 months of my PhD, electron microscopy is indisputably the coolest, most gratifying, and enjoyable. I’m one of those people that loves whatever I’m doing in the moment, as I’m sure my office-mates could attest to. I often come back from trying something new full of enthusiasm and motivation, but that adoration usually dies down after a few days crunching numbers; I end up sitting at my desk pouting over new literature and troubleshooting the protocol. This hasn’t happened with electron microscopy. It has remained steadfast at the front of the pack in the race for my geeky little heart (yes, I actually rank my protocols…don’t we all?), and I don’t see it going anywhere anytime soon.
Over the past few months I have been trained in Transmission Election Microscopy (TEM) and Scanning Electron Microscopy (SEM), and have now completely integrated both into my PhD work. People are usually more familiar with SEM than TEM, but both have their advantages. TEM is commonly used in medical diagnostics as sectioning and staining of different tissues can allow for confirmation and visualisation of various diseases. SEM on the other hand is more often used to assess microstructure and surface detail, such as fractures in building materials. The human eye can only resolve particles that are ~0.2mm apart (for context: the eye of a sewing needle is about 1mm thick), which is small, but not nearly small enough to assess the detail in microscopic organisms. The organisms I work with are only about 10µm wide. These cells are visible through light microscopy but details other than cell shape are not easily discernible.

I use both TEM and SEM for different reasons. I embed, section, and stain Trichodesmium for TEM analyses to visualise intracellular components. TEM resolution is so good that viruses are readily visible, and even macromolecules can be seen under the highest magnifications. Depending on the tissue processed, the preparation for TEM takes about 4 days including preserving, embedding, staining, sectioning, and mounting samples. If you’re really good at sectioning, you can get it all done in 3 days. TEM sections need to be thin (<100nm) because of the way the microscope works. During processing the samples need to be stained with electron dense stains specific to different cell components. That way when the electron beam is focused through the section, scattering occurs in electron dense areas of the section, while areas of the section with little or no staining allow the electrons to pass through. Inside the microscope there is an aperture directly below the specimen, which permits only unscattered electrons through to the next lens; this adds contrast to the image allowing you to discern cellular components. These electrons then pass through a series of lenses and are eventually projected onto a phosphorescent screen that is imaginable. Here, your stained cell components show up dark, and the non-stained portions show up light. Depending on the density of your staining, you can then determine which components are what based not only on shape and size, but also on darkness. This is a very useful tool in my work because the organism I work with is known to undergo intracellular morphological changes before apoptosis (cell death), so I can straightforwardly assess cell integrity with this method.

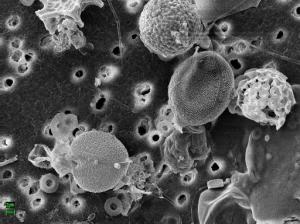
When working with the SEM, I am more interested in outward cell structure, and more specifically, quantifying diatoms and coccolithophores. Sample prep for this microscope is easier than for TEM. While at sea I filter water onto a polycarbonate filter, dry it in the oven, and store it flat until returning to the UK. On the day of analysis I mount a small bit of filter onto a stub and gold coat the sample prior to imaging to increase conductivity. The SEM works by shooting a beam of electrons through the microscope lenses under vacuum towards the sample. When the beam hits the sample, both electrons and X-rays are ejected from the sample, which are detected and sent to a screen. The image is formed by the combination of detected particles scanned. So, in SEM scattered electrons are detected for imaging, while in TEM unscattered electrons form the image.

One of the only disadvantages to these techniques is that all images are in black and white. Colour can only be added artificially after images are produced. Having said that, I don’t see it as a big disadvantage because, at least for my analyses, colour would be distracting. Plus, who could complain about copious amounts of amazing imagery of the coolest organisms on earth? Not me.
Elizabeth Sargent is currently a PhD student at the National Oceanography Centre, Southampton studying nitrogen fixation and its role in fluxes of carbon and nitrogen to the deep sea. She is also a regular contributor for Words in mOcean.
I’m rather glad in my methodology I can simplify the little beasts down to layered spheres 🙂 Epic pictures though! Geeky coolness!
Pretty! This has been an extremely wonderful article.
Thank you for supplying this info.